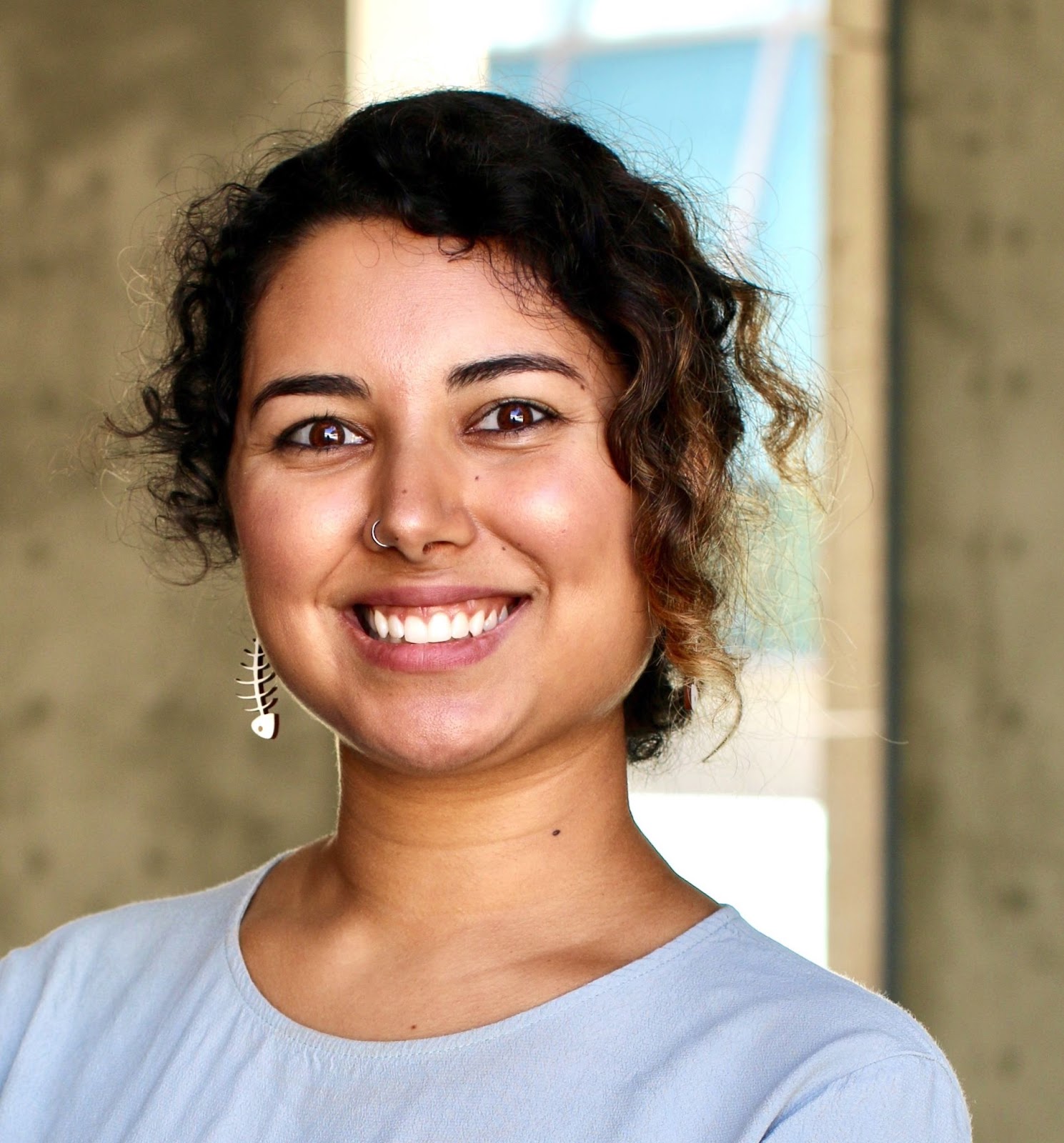
Dr Ul-Hasan (they/them) is a postdoc and lecturer in bioinformatics under Dr Andrew Su and Dr Dawn Eastmond at Scripps Research, doing biocuration and automated data integration work within the Gene Wiki project of Wikidata. They received their PhD in Quantitative & Systems Biology from UC Merced, their Master’s in Biochemistry from the University of New Hampshire and their BSc degrees (3 majors! Biology, Chemistry, and Environmental & Sustainability Studies) from the University of Utah. Sabah is involved in what feels like a thousand different activities related to science, research, coding, outreach, conservation, environmental justice and other things.
I got to know Sabah a couple of years ago when I visited UC Merced and then started following them on twitter. One thing I really love about them is how they don’t limit themselves to just doing one thing.They are ambitious and radical. They founded the Biota project to connect underrepresented communities with nature. They are a filmmaker (see here)! They volunteer for The Carpentries, and they started the venom-microbiome research consortium. They organize workshops, speak at events, teach classes and do many other things.
In my opinion, too few scientists use their platform to fight for justice and to share their passion and knowledge. At the same time, many PhD students and postdocs and even assistant professors are shy about taking a stance, thinking that they would speak up louder (about science or justice or both) when they are more senior. But Sabah proves that you don’t have to be a tenured professor to make a difference in science (they have more than 8000 followers on twitter, just saying).
Pleuni: Hi Sabah, thanks for taking the time to answer my questions! Could you tell us in a few sentences how you became interested in data science?
Sabah: One of my dissertation chapters involved data that was over 100 years old. I know this isn’t a new concept for anyone doing paleo research. I was also well-familiar with “old” data through all the climate change reports that have come up in the public over the years.
However, to directly work with data like that I realized there were so many more questions I wanted to ask people from 100 years ago. That then got me wondering, “How can I contribute to research in a way that can be sustainable 20, 50, or even 5 years from now?”.
My interest in data science thus came from a position of wanting to be part of something bigger in terms of the infrastructure for how we can sustain the science of today and tomorrow.
Pleuni: How did you start learning coding skills? Was it hard for you to learn?
Sabah: I was first introduced to R during my (Biochemistry) Master’s at the University of New Hampshire in 2013. I sat-in on a casual meeting among graduate students and postdocs and truly had no idea what anyone was talking about.
The data analysis section of my MSc thesis ended up utilizing Excel to make bar charts. In retrospect, I see how much faster I could’ve done the analyses if I took the time to learn coding. When I began the doctoral program at UC Merced in January 2015, I knew coding was a skill I wanted to learn and so I did through classes and workshops.
Now it’s my job as a postdoctoral scholar and lecturer for bioinformatics, and I still sometimes struggle with basic concepts. The difference between then and now is I’m a lot better at admitting when I don’t know something, how to ask a question for what I need to learn, and where to go to find that answer.
I’m not sure anyone who does bioinformatics considers themselves an expert, but perhaps the expertise lies within the ability to problem solve especially when it is difficult or can feel overwhelming. In sum, the sooner you can confront your fears the better! Don’t let them freeze you. Believe in your ability to constantly learn and grow, even when you’re a titled expert!
Pleuni: For your paper that appeared in Plos One in 2019, you studied the diversity of microorganisms (including archaea, bacteria and eukaryotes) in seawater and sediment in three different locations. It sounds like a complex dataset to work with.
Community ecology across bacteria, archaea and microbial eukaryotes in the sediment and seawater of coastal Puerto Nuevo, Baja California
Sabah: It’s funny to only be two years out from that publication and already think of so many things I would’ve done differently. I guess that’s growth!
I attribute a lot of credit and thanks to the co-authors of the paper and those in the acknowledgements. It came a long way from when I first drafted it to the final publication form, and posting it on bioRxiv also helped a great deal in soliciting feedback.
What I think really makes a difference is the transparency of that research and associated code, especially in reference to data clean-up (which is the bulk of the analysis work, in personal opinion). I’ve since received several inquiries from people for their own work and to me that feels great to know that it can serve as something people can apply to their own research in making things a little easier.
I also think it’s important we as scientists specify the microbes we’re investigating in any ‘microbial community’ -type paper. Many of the amplicon and metagenomics studies I see really focus on bacteria or fungi, which is absolutely fine but that isn’t a comprehensive microbial community for what many of the titles for these papers tend to imply. In this study, too, we focus on whatever microbial groups we identified solely through 16S and 18S. We need to be better at saying what the data is rather than wordsmithing for a nice story. That will help the next group build upon those gaps for something stronger next time, and overall our intent as scientists is to always have research be advancing further and further. Right?
Pleuni: You used R for your data analysis (but also other software such as QIIME2). What do you like or not like about R? Could you imagine doing a paper like this one without R?
Sabah: Using wrappers such as QIIME2 and mothur are great for people who want to do an analysis of a microbial dataset and then perhaps never touch one again. For me, I found myself continuously asking a lot of “Why?” and wanting to dig deeper on the fundamentals behind what the software I was using. In the end, R took more time to learn short-term but made more sense to me of what was happening each step of the way in the analysis. It was also a good way to affirm my results in trying different avenues and seeing the same output.
What I learned from putting together the paper is it’s not about finding the ‘right’ or ‘wrong’ answer, it’s about finding an answer that is logical and as unbiased as possible. A lot of the time we have these hypotheses we ‘prove’ through confirmation bias. To me, code (when done with intention) is a way to step outside of ourselves and see what the data is telling us rather than what we want the data to say — and that’s where the interesting science lives.
This publication, for example, wasn’t exactly what we were wanting to see. It’s actually a failed attempt at sequencing the venom microbial community of Californiconus californicus, which was the focus of my dissertation (venom microbiomes), due to too much host contamination of the tissues we sampled for that region of Puerto Nuevo. So, what do we do? Do we call it all a wash? There was a lot of thought, time, and resources that went into that work.
I had sampled the sediment and water of the area, along with some generic chemistry tests, to see if the venom microbial community was largely specialized to the snail venom glands or from the surrounding environment (they burrow in the sand). That data was still usable, had good replication, and we didn’t know anything about the microbial community of Puerto Nuevo before that point. Ah-ha! A different story than we were thinking, but still a valuable one. Let the data tell you, don’t misconstrue the data to fit your narrative.
R, and all the programming languages I’ve learned thus far, have helped me learn that.
Pleuni: On your twitter profile, you list many interests, such as advocacy, consulting, data visualization. Can you tell us a bit about your different interests? Are these things linked to each other?
Sabah: Well… haha. The link is that, at heart, I’m a bit of a troublemaker. It’s the nature of a scientist to ask a lot of questions, and asking too many questions can often get us into trouble! I likewise enjoy being asked a lot of questions, and hope to always maintain humility in learning just as much from high school students as I do from tenured professors.
I wanted my Twitter profile and bio to emulate that duality of being both a ‘credible academic’ while also pushing back on what we define as ‘the norm’. I disagree with the idea that a science expert needs to possess a PhD (or some other form of higher education certification) because of the privilege and whiteness involved, but I do also benefit from it after completing the process and there is of course also danger in believing ‘just anyone’ on the internet. And I love learning and helping, which are really the only drivers behind all my many interests.
In my view, the most important quality in being a scientist is being approachable. If only a few people can understand the work you do, then what’s the point? That’s why I’m on Twitter, and also as a way to keep myself grounded, especially learning from moments of being called out (which does happen from time to time). I’d also say my family keeps me in check, as I’m one of the few with a science background. I have one cousin on my Mom’s side with a Ph.D. and that’s it for our extended family of over 100 people (South Asian families are big). Being a good scientist is just as much about humanity as it is about the basic research. I think only good things can come from staying tuned into the reality of the world around us, even though it can feel like a lot to balance.
Pleuni: Do you have any advice for the bio and chem Master’s students in my Data Science class?
Sabah: My advice is to just go for it!
This past Fall I taught a bioinformatics course to (mainly) graduate students and it was an adventure for all of us. It was my first time as a full instructor for a course (versus a teaching assistant), during COVID no less, and it was also the first time many students in the course were getting into bioinformatics.
At the end, it was clear to me that student progress in the course wasn’t about who knew how much at the start but rather about showing up with enthusiasm and simply trying. That went both ways for me as the instructor giving lectures my all as well as for the students and their performance. And life happens! I had to cancel one of the days due to personal life things, and that’s okay. Be good to yourself when you need to and also don’t hold yourself back. And be good to others, too. We really never know what someone else may be experiencing behind the scenes for them to be flakey or on edge, and the more we can find the good in each other the better we can focus on doing the good science.
On that note, I can’t express enough how much of a difference it’s made in my life to work for or alongside with even just one considerate person. As they say, “You are what you eat.”. My PhD co-advisors (Dr Tanja Woyke and Dr Clarissa Nobile) and my current PIs (Dr Su and Dr Eastmond) are truly outstanding people. They have so many stresses in their own careers and lives, and they still somehow show up with kindness and professionalism every day. And they also believe in me to do good work, even when I’ve had a bad week (or month!). That trust really goes such a long way when you’re underrepresented in your field, and often used to being discouraged and/or people expecting very little of you. Being entrusted to teach a course at a renowned research institute directly out of my PhD, for instance, is a big reason why I chose this position in knowing that my voice was heard and respected. That’s been true throughout, and makes it much easier to show up with my best foot forward even on the tough days.
Tying it all together, so many times I’ve got myself stuck because I see others who are ahead of me, doing better than me, and/or with access to more resources than me. One truth we can all agree upon is that life is unfair, and while hopefully it will become equitable over time through our own efforts to create change the fact is that life is still happening in the meantime. No one will help you as much as you can help yourself, and the moments where I’ve been able to just sit down and see something through is how I’ve realized more and more just how much more ability I have than I thought. You’re much more capable than you give yourself credit! It’s super cheesy, but it’s very true. And feel free to reach out any time!
Pleuni: Thanks for answering my questions, Sabah! So much here that resonates with me, including one of the last things you said, that you realized that you have more ability than you thought. This happens to me too! As just one example, just over a year ago, I didn’t think I could learn Machine Learning, but now I am even teaching it. Not that I am suddenly an expert, but I can do it and it is no longer scary.
I look forward to seeing all the science, art, and justice-related projects you will be doing in the future!
Links
Sabah Ul-Hasan Google Scholar profile
Sabah Ul-Hasan, PhD Twitter Profile (@sabahzero)
Tags: bioinformatics, coding, environmental justice, science, scientist spotlight













